I’m a little late due to travel, etc. But here’s the digelst from last Thursday’s final conference presentations.
Sjors Scheres showed the next step in DNA origami enhancement of cryo-EM imaging and tomography. Aleksei Aksimentiev talked about nanopore work.
Zorana Zeravcic was of particular interest to me because she was looking for rule-sets for computational self-assembly of particles. This basically is the theoretical framework for building larger scale structures from nano-powered micromachines. I have been thinking about how to program a microparticle in a “swarm” to act like a state machine, and I have some ideas. I have understood that such machines can be programmed since Eric Klavins’ class in graduate school. But this was another step that I have only a glimmer of an idea about: colloids in 3D creating self-replicating meta-patterns.
Andrew Phillips is from Microsoft. He talked about his meta-language for compiling DNA circuits from a higher level of abstraction. I think that’s great but I have not learned to use his software effectively. I need to learn it and I would love to find/create an index of the software ecosystem around DNA strand escange.
Alexander Deiters talked about his “side projects” in logic gates functioning in human cells. In particular, he made caged DNA that only does strand displacement after UV irradiation and could show that these SD reactions would occur in HEK cells after being transfected in directly. I think that surprised everyone, even him.
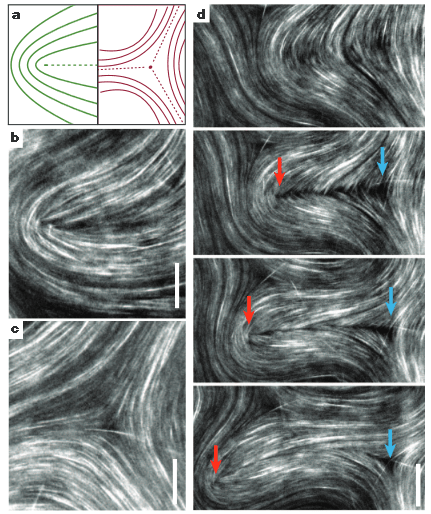
Zev Gartner showed some simply amazing work on using hydrophobic-modified oligonucleotides to label and assembly cells. He makes microtissues that display some amazing emergent behaviors that are not evidently seen in the cell types by themselves. One of the most striking examples was when he put few RAS mutant cells in with many non-mutant cells, he got hyper-motility. But in the reverse scenario there was not hyper motility. Something about the physical/chemical environlemnt of normal cells made the RAS mutant cell behave very interestingly.
Image from: Liu JS, Farlow JT, Paulson AK, Labarge MA, Gartner ZJ. Programmed Cell-to-Cell Variability in Ras Activity Triggers Emergent Behaviors during Mammary Epithelial Morphogenesis. Cell Reports. 2012 Nov 29;2(5):1461–70.



You must be logged in to post a comment.